OUR VISION
From a mechanistic point of view cells can be seen as integrated devices composed of a network of interactions among different macromolecules and small molecules (proteins, nucleic acids and metabolites). Most of them are protein molecular machines that carry out specific tasks, such as building structures and catalyse reactions. In our team we try to understand these complex networks and how proteins are activated. This is also important to design strategies to correct these processes when they get out of control in diseases.
During the last decade a new discipline of biology called proteomics has changed the paradigm of the methods used to study biological mechanisms. Rather than recapitulating the (bio)chemistry of single reactions reconstructed in a test tube, proteomics aims to capture a snapshot of all biological processes and molecular event occurring in vivo by profiling the levels of proteins present in the cell. Unfortunately, knowing protein abundances is not often sufficient to explain phenotypes. This is because the activity of all proteins depends not only on their abundance but also on their 3D structure. Ultimately, all factors that can modify protein’s shapes, most relevantly molecular interactions with metabolites, drugs. nucleic acids and other proteins are important for their function. Thus, a systematic and comprehensive understanding of biological processes has to take into account all these heterogenous interactions.
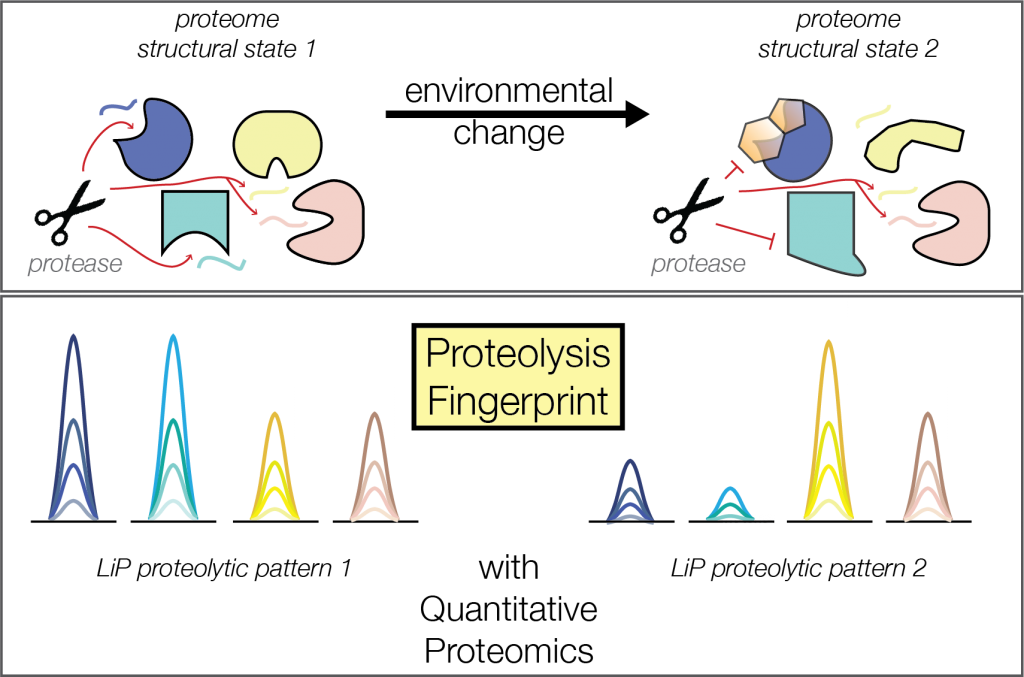
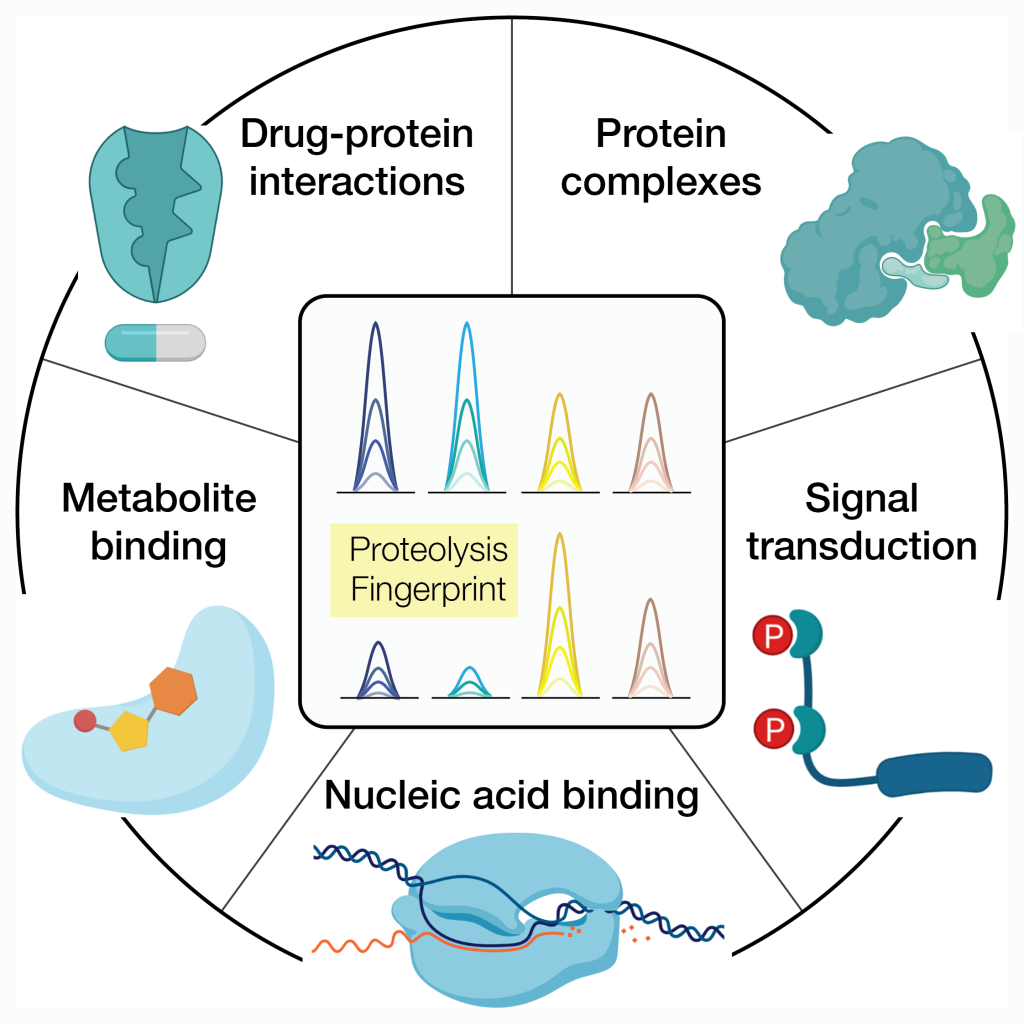
Guided by the paradigm that a change in activity generally correspond to a change in structure, this powerful techniques allows us to study protein structural changes associated to many different types of molecular cues. For instance:
- Assembly-disassembly of protein complexes
- The effects of phosphorylation on protein function
- Interactions of protein to metabolites
- Identification of protein drug targets in vitro and in vivo
- Dynamic interactions of proteins with nucleic acids
- Protein stability changes due to mutations or aggregation
We are also very interested in advancing further the field of structural systems biology by improving new structural proteomics workflows and their applications to different interactomics workflow. With the multidisciplinary research and collaborative atmosphere of the MDC / BIMSB, we aim to combine these state-of-the-art proteomics methods with genomics, metabolomics, stem cell biology and computational biology. Our general goal Is to advance the general knowledge of basic research and make a direct impact on topics related to human health. We are in the right environment to do so.
